Routinely testing your R package with Travis
This protocol was contributed by L. Collado-Torres.
Overview
At some point after in the R (R Core Team, 2015) learning curve you get to a level where you want to write your own functions. The next step is to share the functions in the form of an R package. The Leek group way of doing so is described in more detail here (jtleek, 2015).
Sharing your functions as an R package means that others can easily use them after installing the package. This all sounds great until you then realize that there is some responsibility on your side to make sure things are working properly. R has a command to help you do so, namely R CMD check which verifies that everything is in order.
Projects like Bioconductor (Huber, W., Carey, J., et al., 2015) help both the developer and the user community by running R CMD check on their packages every day. Doing so helps identify any issues your functions might have because something else changed (like a function you imported). Bioconductor runs other tests and demands that a package satisfies specific criteria to be included in the project, but the idea of routinely checking your R package remains.
In recent years, more and more developers have turned towards hosting their packages in GitHub, specially in the development phase even when working with CRAN or Bioconductor (through a git-svn bridge). While as a developer you can test your package locally, at some point it starts to take a large chunk of your time. That's where open source tools like Travis CI make your life as a developer easier. You can basically set Travis CI to run R CMD check for you every time you push code to GitHub. This protocol explains how to setup Travis CI.
Setting up Travis
To start, we will assume that you have a GitHub account with at least one repository that contains an R package, lets call it rpack. Your very first step is to sign in to Travis using your GitHub account at travis-ci.org/.
Basic setup
Flip the switch
Once Travis gets your list of GitHub repositories (force a sync if you don't see rpack), you can turn on Travis for rpack by flipping the switch at travis-ci.org/profile. For example, here we turned on the switch for derfinder:
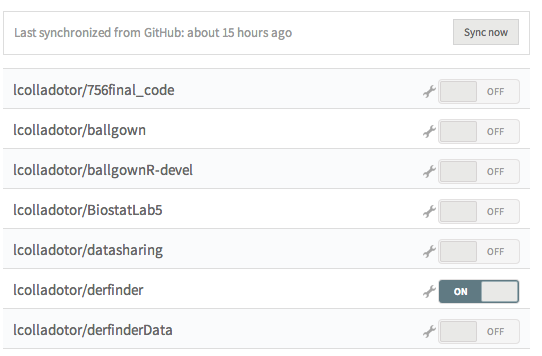
Setup via devtools
If you are using devtools (Wickham and Chang, 2015) for developing your package, you can simply add Travis to rpack by using
library('devtools')
use_travis('path to rpack')This will create the .travis.yml and .Rbuildignore files in rpack's main directory.
Manual basic setup
Add .travis.yml
The next step is to add the file .travis.yml to your repository (in the main directory). This file will specify your Travis configuration such as which R packages to install, when you want to be notified by Travis, and what information you want to see.
The easiest way to setup Travis for R projects is to use r-travis (craigcitro, 2015). r-travis instructions show how to add .travis.yml with commands, but basically, you want to start by copying the example configuration file shown below into your .travis.yml file:
# Sample .travis.yml for R projects.
#
# See README.md for instructions, or for more configuration options,
# see the wiki:
# https://github.com/craigcitro/r-travis/wiki
language: c
before_install:
- curl -OL http://raw.github.com/craigcitro/r-travis/master/scripts/travis-tool.sh
- chmod 755 ./travis-tool.sh
- ./travis-tool.sh bootstrap
install:
- ./travis-tool.sh install_deps
script: ./travis-tool.sh run_tests
after_failure:
- ./travis-tool.sh dump_logs
notifications:
email:
on_success: change
on_failure: change(Check here for the latest version of this example configuration.)
Ignore .travis.yml
Once you added the .travis.yml file you are nearly done with the initial setup. Next, you have to tell R to ignore this specific file. You can do so using the hidden file .Rbuildignore which should look like this:
## Ignore travis config file
^\.travis\.yml$Add them to git
The last step in the basic setup is to simply ask Git to version control the .travis.yml and .Rbuildignore files. That is, use git add or the add button if you are using a Git/GitHub GUI.
git add .travis.yml .Rbuildignore
git commit -m 'enable continuous integration via craigcitro/r-travis'
git pushThen, push the files to GitHub. This will prompt Travis to check your package using the basic configuration file.
Add a status image
If you want users to quickly check whether your package is passing the tests run on Travis, simply add a status image. Doing so is quite strait forward and explained in more detail here.
Configuring .travis.yml
Installing required packages
On a simple scenario, you only have to use the basic setup. In a more realistic scenario, your package might depend on other packages which are available either through CRAN, Bioconductor or GitHub. If that's the case, r-travis has commands which allow you to install these dependencies before installing rpack.
These are:
install:
## For a CRAN package
- ./travis-tool.sh install_r <package>
## For a Bioconductor package
- ./travis-tool.sh install_bioc <package>
## For a GitHub package
- ./travis-tool.sh install_github <package>These can be useful if you want to install the packages in a specific order. However, you might prefer to use commands that look up rpack's DESCRIPTION file and automatically determine the list of packages to install.
install:
## For installing all CRAN dependencies using rpack's DESCRIPTION
- ./travis-tool.sh install_deps
## For installing all Bioconductor dependencies using rpack's DESCRIPTION
- ./travis-tool.sh install_bioc_depsAny of these commands have to be specified under the install step. For example
install:
- ./travis-tool.sh install_github hadley/devtools
- ./travis-tool.sh install_bioc_depsNote that if you are installing Bioconductor packages, there are two versions you could be using. The current release or the current devel version. By default, r-travis uses the current devel version of the packages, but you can turn this off by specifying in .travis.yml the BIOC_USE_DEVEL environment variable:
env:
global:
- BIOC_USE_DEVEL="FALSE"Further configuration details are explained in the r-travis wiki. You might also be interested in checking the travis-examples repository.
Configuring R CMD check
By default r-travis will skip the step of building the vignette and the manual in R CMD build and R CMD check. For checking, it will also test as it was for CRAN. More specifically, the default options are:
R CMD build --no-build-vignettes --no-manual rpack
R CMD check --no-build-vignettes --no-manual --as-cran rpackIn your case, you might want to use a different set of options to build and/or check your package. For example, you might be interested in getting the timing information for the examples for all of your functions. You can do so by setting the environment variable _R_CHECK_TIMINGS_ to 0.
That is, using the following in your .travis.yml
env:
global:
- R_CHECK_ARGS="--no-build-vignettes --no-manual --timings"
- _R_CHECK_TIMINGS_="0"_R_CHECK_TIMINGS_ and other options are described in more detail at the R Internals manual.
Configuring the report
Checking for how much time was spent in each of the example pages is great (see above), but by default this information is not reported on Travis. To get the timing information you need to configure Travis to dump the data in the report.
Similarly, you might want Travis to report the versions of the R packages used in the test. To achieve both goals, add the following to your .travis.yml file:
after_script:
- ./travis-tool.sh dump_logs_by_extension "timings"
- ./travis-tool.sh dump_sysinfoderfinder use case
As a complicated use case, we present the .travis.yml file for the derfinder package.
derfinder depends on two other packages that are available via GitHub: derfinderHelper and derfinderData. Thus, they need to be installed before derfinder can be installed. However, those two packages have some dependencies themselves such as knitr and rmarkdown, so they have to be installed first.
Furthermore, derfinder runs its tests only then the environment variable R_CHECK_TESTS is set to TRUE (details here). Finally, the results from the Travis tests are reported to Slack: for details check the Slack-Travis integration site.
language: c
before_install:
# - curl -OL http://raw.github.com/craigcitro/r-travis/master/scripts/travis-tool.sh
- curl -OL http://raw.github.com/lcolladotor/r-travis/master/scripts/travis-tool.sh
- chmod 755 ./travis-tool.sh
- ./travis-tool.sh bootstrap
install:
- ./travis-tool.sh install_bioc S4Vectors
- ./travis-tool.sh install_bioc IRanges
- ./travis-tool.sh install_r Matrix
- ./travis-tool.sh install_r knitr
- ./travis-tool.sh install_r rmarkdown
- ./travis-tool.sh install_r knitcitations
- ./travis-tool.sh install_r knitrBootstrap
- ./travis-tool.sh install_github lcolladotor/derfinderHelper
- ./travis-tool.sh install_github lcolladotor/derfinderData
- ./travis-tool.sh install_bioc_deps
script: ./travis-tool.sh run_tests
on_failure:
- ./travis-tool.sh dump_logs
after_script:
- ./travis-tool.sh dump_logs_by_extension "timings"
- ./travis-tool.sh dump_sysinfo
notifications:
email:
on_success: change
on_failure: change
slack:
secure: FIA40TI4UkOHvR19rNCfX1la5tiCmyEMjiO/sGyK0cWGt5qQxIOp+PHE3pOk9axYiVacbSCR3oAosQUsOxRew/6FyMsNR3bCPXVUzrIimABvBbjofMBDx3Z7W03O+6YahmHwDrJEWDuJ0k4457QqeqhITcFWX4twieo5fJCjedI=
env:
global:
- R_BUILD_ARGS="--no-build-vignettes --no-manual --no-resave-data"
- R_CHECK_ARGS="--no-build-vignettes --no-manual --timings"
- R_CHECK_TIME="TRUE"
- R_CHECK_TESTS="TRUE"
- _R_CHECK_TIMINGS_="0"(The latest version can be seen live here)
Learn more
If you already browsed the r-travis wiki the next best place is to actually look at travis-tool.sh. This is the main script that runs r-travis.
References
We first heard about Travis CI integration with R from Yihui Xie on his blog post Travis CI for R? and it's follow up post Travis CI for R? (not yet). However, it only became easy to install thanks to the r-travis project and with [@siddharthab](https://github.com/siddharthab)'s contributions to support Bioconductor packages. Plus a helpful reminder to take a look at Travis CI:
Are you building packages for #rstats, using @github, and not using @travisci? Stop and look at devtools::add_travis now.
— John Muschelli (@StrictlyStat) September 11, 2014
Citations made with knitcitations (Boettiger, 2015).
[1] C. Boettiger. knitcitations: Citations for Knitr Markdown Files. R package version 1.0.6. 2015. URL: http://CRAN.R-project.org/package=knitcitations.
[2] craigcitro. craigcitro/r-travis. https://github.com/craigcitro/r-travis. 2015. URL: https://github.com/craigcitro/r-travis.
[3] Huber, W., Carey, V. J., et al. “Orchestrating high-throughput genomic analysis with Bioconductor”. In: Nature Methods 12.2 (2015), pp. 115–121. URL: http://www.nature.com/nmeth/journal/v12/n2/full/nmeth.3252.html.
[4] jtleek. jtleek/rpackages. https://github.com/jtleek/rpackages. 2015. URL: https://github.com/jtleek/rpackages.
[5] R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. Vienna, Austria, 2015. URL: http://www.R-project.org/.
[6] H. Wickham and W. Chang. devtools: Tools to Make Developing R Packages Easier. R package version 1.8.0. 2015. URL: http://CRAN.R-project.org/package=devtools.